Conserved signature indels
Conserved signature inserts and deletions (CSIs) in protein sequences provide an important category of molecular markers for understanding phylogenetic relationships.[1][2] CSIs, brought about by rare genetic changes, provide useful phylogenetic markers that are generally of defined size and they are flanked on both sides by conserved regions to ensure their reliability. While indels can be arbitrary inserts or deletions, CSIs are defined as only those protein indels that are present within conserved regions of the protein.[2][3][4][5]
The CSIs that are restricted to a particular clade or group of species, generally provide good phylogenetic markers of common evolutionary descent.[2] Due to the rarity and highly specific nature of such changes, it is less likely that they could arise independently by either convergent or parallel evolution (i.e. homoplasy) and therefore are likely to represent synapomorphy. Other confounding factors such as differences in evolutionary rates at different sites or among different species also generally do not affect the interpretation of a CSI.[2][3] By determining the presence or absence of CSIs in an out-group species, one can infer whether the ancestral form of the CSI was an insert or deletion and this can be used to develop a rooted phylogenetic relationship among organisms.[1][2]
CSIs are discovered by looking for shared changes in a phylogenetic tree constructed from protein sequences. Most CSIs that have been identified have been found to have high predictive value upon addition of new sequences, retaining the specificity for the originally identified clades of species. They can be used to identify both known and even previously unknown species belonging to these groups in different environments.[3] Compared to tree branching orders which can vary among methods, specific CSIs make for more concrete circumscriptions that are computationally cheaper to apply.[6]
Types
[edit]Group-specific
[edit]
Group-specific CSIs are commonly shared by different species belonging to a particular taxon (e.g. genus, family, class, order, phylum) but they are not present in other groups. These CSIs were most likely introduced in an ancestor of the group of species before the members of the taxa diverged. They provide molecular means for distinguishing members of a particular taxon from all other organisms.[2][5]
Figure 1 shows an example of 5aa CSI found in all species belonging to the taxon X. This is a distinctive characteristic of this taxon as it is not found in any other species. This signature was likely introduced in a common ancestor of the species from this taxon. Similarly other group-specific signatures (not shown) could be shared by either A1 and A2 or B1 and B2, etc., or even by X1 and X2 or by X3 and X4, etc. The groups A, B, C, D and X, in this diagram could correspond to various bacterial or Eukaryotic phyla.[7]
Group-specific CSIs have been used in the past to determine the phylogenetic relationship of a number of bacterial phyla and subgroups within it. For example a 3 amino acid insert was uniquely shared by members of the phylum Thermotogota (formerly Thermotogae) in the essential 50S ribosomal protein L7/L12, within a highly conserved region (82-124 amino acid). This is not present in any other bacteria species and could be used to characterize members of Thermotogota from all other bacteria. Group-specific CSIs were also used to characterize subgroups within Thermotogota.[8]
Multi-group or mainline
[edit]
Mainline CSIs are those in which a conserved insert or deletion is shared by several major phyla, but absent from other phyla.[2]
Figure 2 shows an example of 5aa CSI found in a conserved region that is commonly present in the species belonging to phyla X, Y and Z, but it is absent in other phyla (A, B and C). This signature indicates a specific relationship of taxa X, Y and Z and also A, B and C. Based upon the presence or absence of such an indel, in out-group species (viz. Archaea), it can be inferred whether the indel is an insert or a deletion, and which of these two groups A, B, C or X, Y, Z is ancestral.[7]
Mainline CSIs have been used in the past to determine the phylogenetic relationship of a number of bacterial phyla. The large CSI of about 150-180 amino acids within a conserved region of Gyrase B (between amino acids 529-751), is commonly shared between various Pseudomonadota, Chlamydiota, Planctomycetota and Aquificota species. This CSI is absent in other ancestral bacterial phyla as well as Archaea.[9] Similarly a large CSI of about 100 amino acids in RpoB homologs (between amino acids 919-1058) is present in various species belonging to Pseudomonadota, Bacteroidota, Chlorobiota, Chlamydiota, Planctomycetota, and Aquificota. This CSI is absent in other ancestral bacterial phyla as well as Archaea.[10][11] In both cases one can infer that the groups lacking the CSI are ancestral.
Evolutionary studies based on CSIs
[edit]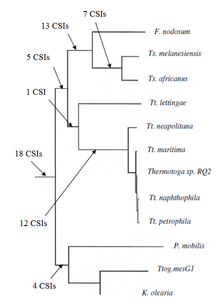


A key issue in bacterial phylogeny is to understand how different bacterial species are related to each other and their branching order from a common ancestor. Currently most phylogenetic trees are based on 16S rRNA or other genes/proteins. These trees are not always able to resolve key phylogenetic questions with a high degree of certainty.[12][13][14][15][16] However in recent years the discovery and analyses of conserved indels (CSIs) in many universally distributed proteins have aided in this quest. The genetic events leading to them are postulated to have occurred at important evolutionary branch points and their species distribution patterns provide valuable information regarding the branching order and interrelationships among different bacterial phyla.[1][2][8]
Thermotogota
[edit]Recently the phylogenetic relationship of the group Thermotogota was characterized based on the CSI approach. Previously no biochemical or molecular markers were known that could clearly distinguish the species of this phylum from all other bacteria. More than 60 CSIs that were specific for the entire Thermotogota phylum or its different subgroups were discovered. Of these, 18 CSIs are uniquely present in various Thermotogota species and provide molecular markers for the phylum. Additionally there were many CSIs that were specific for various Thermotogota subgroups. Another 12 CSIs were specific for a clade consisting of various Thermotogota species except Tt. Lettingae. While 14 CSIs were specific for a clade consisting of the Fervidobacterium and Thermosipho genera and 18 CSIs were specific for the genus Thermosiphon.[citation needed]
Lastly 16 CSIs were reported that were shared by either some or all Thermotogota species or some species from other taxa such as Archaea, Aquificota, Bacillota, Pseudomonadota, Deinococcota, Fusobacteriota, Dictyoglomota, Chloroflexota, and eukaryotes. The shared presence of some of these CSIs could be due to lateral gene transfer (LGT) between these groups. However the number of CSIs that are commonly shared with other taxa is much smaller than those that are specific for Thermotogota and they do not exhibit any specific pattern. Hence they have no significant effect on the distinction of Thermotogota.[8]
Archaea
[edit]Mesophillic Thermoproteota were recently placed into a new phylum of Archaea called the Nitrososphaerota (formerly Thaumarchaeota). However there are very few molecular markers that can distinguish this group of archaea from the phylum Thermoproteota (formerly Crenarchaeota). A detailed phylogenetic study using the CSI approach was conducted to distinguish these phyla in molecular terms. 6 CSIs were uniquely found in various Nitrososphaerota, namely Cenarchaeum symbiosum, Nitrosopumilus maritimus and a number of uncultured marine Thermoproteota. 3 CSIs were found that were commonly shared between species belonging to Nitrososphaerota and Thermoproteota. Additionally, a number of CSIs were found that are specific for different orders of Thermoproteota—3 CSIs for Sulfolobales, 5 CSIs for Thermoproteales, lastly 2 CSIs common for Sulfolobales and Desulfurococcales. The signatures described provide novel means for distinguishing Thermoproteota and Nitrososphaerota, additionally they could be used as a tool for the classification and identification of related species.[17]
Pasteurellales
[edit]The members of the order Pasteurellales are currently distinguished mainly based on their position in the branching of the 16srRNA tree. There are currently very few molecular markers known that can distinguish members of this order from other bacteria. A CSI approach was recently used to elucidate the phylogenetic relationships between the species in this order; more than 40 CSIs were discovered that were uniquely shared by all or most of the species. Two major clades are formed within this Pasteurellales: Clade I, encompassing Aggregatibacter, Pasteurella, Actinobacillus succinogenes, Mannheimia succiniciproducens, Haemophilus influenzae and Haemophilus somnus, was supported by 13 CSIs. Clade II, encompassing Actinobacillus pleuropneumoniae, Actinobacillus minor, Haemophilus ducreyi, Mannheimia haemolytica and Haemophilus parasuis, was supported by 9 CSIs. Based on these results, it was proposed that Pasteurellales be divided from its current one family into two different ones. Additionally, the signatures described would provide novel means of identifying undiscovered Pasteurellales species.[18]
Gammaproteobacteria
[edit]The class Gammaproteobacteria forms one of the largest groups of bacteria. It is currently distinguished from other bacteria solely by 16s rRNA-based phylogenetic trees. No molecular characteristics unique to the class or its different subgroups are known. A detailed CSI-based study was conducted to better understand the phylogeny of this class. Firstly, a phylogenetic tree based on concatenated sequences of a number of universally-distributed proteins was created. The branching order of the different orders of the class Gammaproteobacteria (from most recent to the earliest diverging) was: Enterobacteriales >Pasteurellales >Vibrionales, Aeromonadales >Alteromonadales >Oceanospirillales, Pseudomonadales >Chromatiales, Legionellales, Methylococcales, Xanthomonadales, Cardiobacteriales, Thiotrichales. Additionally, 4 CSIs were discovered that were unique to most species of the class Gammaproteobacteria. A 2 aa deletion in AICAR transformylase was uniquely shared by all gammaproteobacteria except for Francisella tularensis. A 4 aa deletion in RNA polymerase b-subunit and a 1 aa deletion in ribosomal protein L16 were found uniquely in various species belonging to the orders Enterobacteriales, Pasteurellales, Vibrionales, Aeromonadales and Alteromonadales, but were not found in other gammaproteobacteria. Lastly, a 2 aa deletion in leucyl-tRNA synthetase was commonly present in the above orders of the class Gammaproteobacteria and in some members of the order Oceanospirillales.[19] Another CSI-based study has also identified 4 CSIs that are exclusive to the order Xanthomonadales. Taken together, these two facts show that Xanthomonadales is a monophyletic group that is ancestral to other Gammaproteobacteria, which further shows that Xanthomonadales is an independent subdivision, and constitutes one of the deepest-branching lineages within the Gammaproteobacteria clade.[4][19]
See also
[edit]References
[edit]- ^ a b c Baldauf, S. L. (1993). "Animals and Fungi are Each Other's Closest Relatives: Congruent Evidence from Multiple Proteins". Proceedings of the National Academy of Sciences. 90 (24): 11558–11562. Bibcode:1993PNAS...9011558B. doi:10.1073/pnas.90.24.11558. PMC 48023. PMID 8265589.
- ^ a b c d e f g h Gupta, Radhey S. (1998). "Protein Phylogenies and Signature Sequences: A Reappraisal of Evolutionary Relationships among Archaebacteria, Eubacteria, and Eukaryotes". Microbiology and Molecular Biology Reviews. 62 (4): 1435–91. doi:10.1128/MMBR.62.4.1435-1491.1998. PMC 98952. PMID 9841678.
- ^ a b c Gupta, Radhey S.; Griffiths, Emma (2002). "Critical Issues in Bacterial Phylogeny". Theoretical Population Biology. 61 (4): 423–34. doi:10.1006/tpbi.2002.1589. PMID 12167362.
- ^ a b Cutiño-Jiménez, Ania M.; Martins-Pinheiro, Marinalva; Lima, Wanessa C.; Martín-Tornet, Alexander; Morales, Osleidys G.; Menck, Carlos F.M. (2010). "Evolutionary placement of Xanthomonadales based on conserved protein signature sequences". Molecular Phylogenetics and Evolution. 54 (2): 524–34. doi:10.1016/j.ympev.2009.09.026. PMID 19786109.
- ^ a b Rokas, Antonis; Holland, Peter W.H. (2000). "Rare genomic changes as a tool for phylogenetics". Trends in Ecology & Evolution. 15 (11): 454–459. doi:10.1016/S0169-5347(00)01967-4. PMID 11050348.
- ^ Gupta, Radhey S.; Kanter-Eivin, David A. (9 May 2023). "AppIndels.com server: a web-based tool for the identification of known taxon-specific conserved signature indels in genome sequences. Validation of its usefulness by predicting the taxonomic affiliation of >700 unclassified strains of Bacillus species". International Journal of Systematic and Evolutionary Microbiology. 73 (5). doi:10.1099/ijsem.0.005844. PMID 37159410.
- ^ a b Gupta, Radhey. "Conserved Inserts and Deletions in Protein Sequences". Bacterial Phylogeny. Gupta lab. Archived from the original on 15 September 2011. Retrieved 2 April 2012.
{{cite web}}: CS1 maint: bot: original URL status unknown (link) - ^ a b c Gupta, Radhey S.; Bhandari, Vaibhav (2011). "Phylogeny and molecular signatures for the phylum Thermotogae and its subgroups". Antonie van Leeuwenhoek. 100 (1): 1–34. doi:10.1007/s10482-011-9576-z. PMID 21503713. S2CID 24995263.
- ^ Griffiths, E.; Gupta, R. S. (2007). "Phylogeny and shared conserved inserts in proteins provide evidence that Verrucomicrobia are the closest known free-living relatives of chlamydiae". Microbiology. 153 (8): 2648–54. doi:10.1099/mic.0.2007/009118-0. PMID 17660429.
- ^ Gupta, Radhey S. (2003). "Evolutionary relationships among photosynthetic bacteria". Photosynthesis Research. 76 (1–3): 173–83. doi:10.1023/A:1024999314839. PMID 16228576. S2CID 38460308.
- ^ Griffiths, Emma; Gupta, Radhey S. (2004). "Signature sequences in diverse proteins provide evidence for the late divergence of the Order Aquificales" (PDF). International Microbiology. 7 (1): 41–52. PMID 15179606.
- ^ Brown, James R.; Douady, Christophe J.; Italia, Michael J.; Marshall, William E.; Stanhope, Michael J. (2001). "Universal trees based on large combined protein sequence data sets". Nature Genetics. 28 (3): 281–5. doi:10.1038/90129. PMID 11431701. S2CID 8516570.
- ^ Cavalier-Smith, T (2002). "The neomuran origin of archaebacteria, the negibacterial root of the universal tree and bacterial megaclassification". International Journal of Systematic and Evolutionary Microbiology. 52 (1): 7–76. doi:10.1099/00207713-52-1-7. PMID 11837318.
- ^ Ciccarelli, F. D.; Doerks, T; Von Mering, C; Creevey, CJ; Snel, B; Bork, P (2006). "Toward Automatic Reconstruction of a Highly Resolved Tree of Life". Science. 311 (5765): 1283–7. Bibcode:2006Sci...311.1283C. CiteSeerX 10.1.1.381.9514. doi:10.1126/science.1123061. PMID 16513982. S2CID 1615592.
- ^ Daubin, V.; Gouy, M; Perrière, G (2002). "A Phylogenomic Approach to Bacterial Phylogeny: Evidence of a Core of Genes Sharing a Common History". Genome Research. 12 (7): 1080–90. doi:10.1101/gr.187002. PMC 186629. PMID 12097345.
- ^ Eisen, Jonathan A. (1995). "The RecA Protein as a Model Molecule for Molecular Systematic Studies of Bacteria: Comparison of Trees of RecAs and 16S rRNAs from the Same Species". Journal of Molecular Evolution. 41 (6): 1105–23. Bibcode:1995JMolE..41.1105E. doi:10.1007/bf00173192. PMC 3188426. PMID 8587109.
- ^ Gupta, Radhey S.; Shami, Ali (2010). "Molecular signatures for the Crenarchaeota and the Thaumarchaeota". Antonie van Leeuwenhoek. 99 (2): 133–57. doi:10.1007/s10482-010-9488-3. PMID 20711675. S2CID 12874800.
- ^ Naushad, Hafiz Sohail; Gupta, Radhey S. (2011). "Molecular signatures (conserved indels) in protein sequences that are specific for the order Pasteurellales and distinguish two of its main clades". Antonie van Leeuwenhoek. 101 (1): 105–24. doi:10.1007/s10482-011-9628-4. PMID 21830122. S2CID 15114511.
- ^ a b Gao, B.; Mohan, R.; Gupta, R. S. (2009). "Phylogenomics and protein signatures elucidating the evolutionary relationships among the Gammaproteobacteria". International Journal of Systematic and Evolutionary Microbiology. 59 (2): 234–47. doi:10.1099/ijs.0.002741-0. PMID 19196760.
